Nanouniverse: Virtual Instancing of Structural Detail and Adaptive Shell Mapping
Authors
Ruwayda Alharbi, Ondřej Strnad, Markus Hadwiger, Ivan ViolaDescription
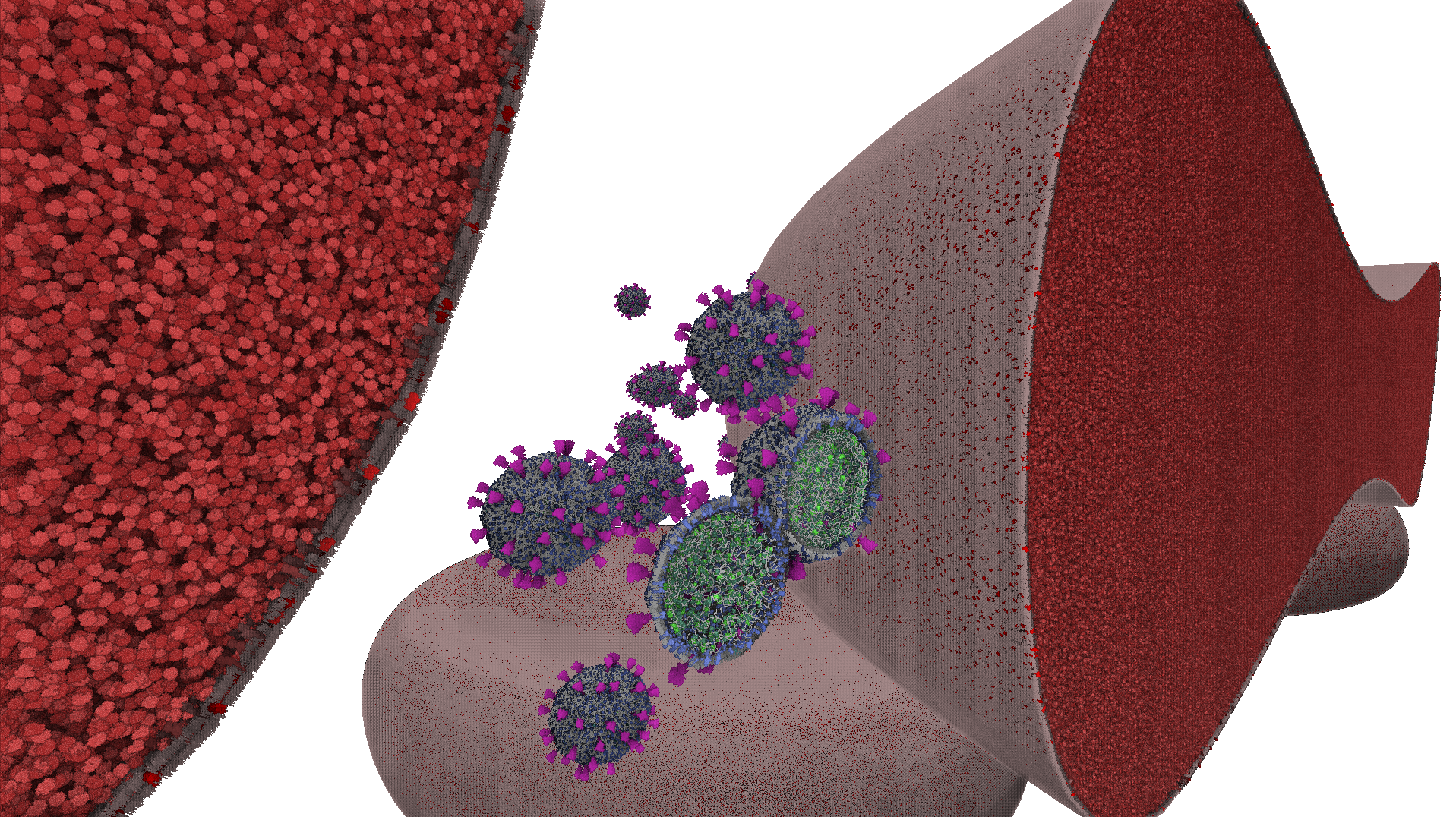
Rendering huge biological scenes with atomistic detail presents a significant challenge in molecular visualization due to the memory limitations inherent in traditional rendering approaches. In this paper, we propose a novel method for the interactive rendering of massive molecular scenes based on hardware-accelerated ray tracing. Our approach circumvents GPU memory constraints by introducing virtual instantiation of full-detail scene elements. Using instancing significantly reduces memory consumption while preserving the full atomistic detail of scenes comprising trillions of atoms, with interactive rendering performance and completely free user exploration. We utilize coarse meshes as proxy geometries to approximate the overall shape of biological compartments, and access all atomistic detail dynamically during ray tracing. We do this via a novel adaptive technique utilizing a volumetric shell layer of prisms extruded around proxy geometry triangles, and a virtual volume grid for the interior of each compartment. Our algorithm scales to enormous molecular scenes with minimal memory consumption and the potential to accommodate even larger scenes. Our method also supports advanced effects such as clipping planes and animations. We demonstrate the efficiency and scalability of our approach by rendering tens of instances of Red Blood Cell and SARS-CoV-2 models theoretically containing more than 20 trillion atoms.